Technical Documentation
The downscaling workflow predicts carbon pools (Soil Organic Carbon and Aboveground Biomass) for cropland fields in California and then aggregates these predictions to the county scale.
It uses an ensemble-based approach to uncertainty propagation and analysis, maintaining ensemble structure to propagate errors through the prediction and aggregation processes.
Terminology
- Design Points: Fields chosen via stratified random sampling using k-means clustering on environmental data layers across California crop fields.
- Crop Fields: All croplands in the LandIQ dataset.
- Anchor Sites: Sites used as ground truth for calibration and validation, including UC research stations and Ameriflux sites.
This Repository Contains Two Workflows That Will Be Split
TODO Workflows (1 & 3) are in this repository and will be split
The workflows are
- Site Selection: uses environmental variables (later also management layers) to create clusters and then select representative sites. The design_points.csv are then passed to the ensemble workflow
- Ensemble in ccmmf/workflows repository, generates ensemble outputs
- Downscaling: uses ensemble outputs to make predictions for each field in CA then aggregate to county level summaries.
Workflow Steps
Configuration
Workflow settings are configured in 000-config.R.
The configuration script reads the CCMMF directory from the environment variable CCMMF_DIR (set in .Renviron), and uses it to define paths for inputs and outputs.
The CCMMF_DIR variable, however, is defined in .Renviron so that it can: - Be used to locate and override R library (RENV_PATHS_LIBRARY) and cache (RENV_PATHS_CACHE) paths outside the home directory. - Be easily overridden via a shell export (export CCMMF_DIR=…) without modifying workflow scripts.
Configuration setup
To set up this workflow to run on your system, follow the following steps.
Clone Repository
git clone git@github.com:ccmmf/downscalingCCMMF_DIRshould point to the shared CCMMF directory. This is the location where data, inputs, and outputs will be stored, as well as the location of therenvcache and libraryRENV_PATHS_CACHEandRENV_PATHS_LIBRARYstore therenvcache and library in the CCMMF directory. These are in a subdirectory of the CCMMF directory in order to make them available across all users (and because on some computers, they exceed allocated space in the home directory).R_LIBS_USERmust point to the platform and R version specific subdirectory insideRENV_PATHS_LIBRARY.
.Rprofile- sets repositories from which R packages are installed
- runs
renv/activate.R
000-config.R- set
pecan_outdirbased on the CCMMF_DIR. - confirm that relative paths (
data_raw,data,cache) are correct. - detect and use resources for parallel processing (with future package); default is
available cores - 1 - PRODUCTION mode setting. For testing, set
PRODUCTIONtoFALSE. This is much faster and requires fewer computing resources because it subsets large datasets. Once a test run is successful, setPRODUCTIONtoTRUEto run the full workflow.
- set
Management Scenario Configuration
To compare carbon outcomes under different agricultural practices, enable management scenario mode:
USE_PHASE_3_SCENARIOS <- TRUEWhen enabled, the workflow processes multiple management scenarios in a single run. Each scenario represents a different combination of practices applied to annual croplands.
Available Management Scenarios:
| Scenario | Description |
|---|---|
| baseline | Conventional management practices |
| compost | Compost application replacing mineral N fertilizer |
| reduced_till | Reduced tillage intensity (0.10) |
| zero_till | No tillage (0.00) |
| reduced_irrig_drip | Drip irrigation replacing canopy irrigation |
| stacked | Combined: compost + reduced tillage + drip irrigation |
1. Data Preparation
Rscript scripts/009_update_landiq.R
Rscript scripts/010_prepare_covariates.R
Rscript scripts/011_prepare_anchor_sites.RThese scripts prepare data for clustering and downscaling:
- Converts LandIQ-derived shapefiles to a geopackage with geospatial information and a CSV with other attributes
- Extracts environmental covariates (clay, organic carbon, topographic wetness, temperature, precipitation, solar radiation, vapor pressure)
- Groups fields into Cal-Adapt climate regions
- Assigns anchor sites to fields
Inputs:
- LandIQ Crop Map:
data_raw/i15_Crop_Mapping_2016_SHP/i15_Crop_Mapping_2016.shp - Soilgrids:
clay_0-5cm_mean.tifandocd_0-5cm_mean.tifConsider aggregating 0-5,5-15,15-30 into a single 0-30 cm layer - TWI:
TWI/TWI_resample.tiff - ERA5 Met Data: Files in
GridMET/folder namedERA5_met_<YYYY>.tiff - Anchor Sites:
data_raw/anchor_sites.csv
Outputs:
ca_fields.gpkg: Spatial information from LandIQca_field_attributes.csv: Site attributes including crop typesite_covariates.csv: Environmental covariates for each fieldanchor_sites_ids.csv: Anchor site information
Environmental Covariates
| Variable | Description | Source | Units |
|---|---|---|---|
| temp | Mean annual temperature | ERA5 | °C |
| precip | Mean annual precipitation | ERA5 | mm/year |
| srad | Solar radiation | ERA5 | W/m² |
| vapr | Vapor pressure deficit | ERA5 | kPa |
| clay | Clay content | SoilGrids | % |
| ocd | Organic carbon density | SoilGrids | g/kg |
| twi | Topographic wetness index | SRTM-derived | - |
2. Design Point Selection
Rscript scripts/020_cluster_and_select_design_points.R
Rscript scripts/021_clustering_diagnostics.RUses k-means clustering to select representative fields plus anchor sites:
- Subsample LandIQ fields and include anchor sites for clustering
- Select cluster number based on the Elbow Method
- Cluster fields using k-means based on environmental covariates
- Select design points from clusters for SIPNET simulation
Inputs: - data/site_covariates.csv - data/anchor_sites_ids.csv
Output: - data/design_points.csv
3. SIPNET Model Runs
A separate workflow prepares inputs and runs SIPNET simulations for the design points.
Inputs: - design_points.csv - Initial conditions (from modeling workflow)
Outputs: - out/ENS-<ensemble_number>-<site_id>/YYYY.nc: NetCDF files containing SIPNET outputs, in PEcAn standard model output format. - out/ENS-<ensemble_number>-<site_id>/YYYY.nc.var: List of variables included in SIPNET output (see table below)
Available Variables
Each output file named YYYY.nc has an associated file named YYYY.nc.var. This file contains a list of variables included in the output. SIPNET outputs have been converted to PEcAn standard units and stored in PEcAn standard NetCDF files. PEcAn standard units are SI, following the Climate Forecasting standards:
- Mass pools: kg / m2
- TotSoilCarb: Total Soil Carbon
- AbvGrndWood: Above ground woody biomass
- Mass fluxes: kg / m2 / s-1
- GPP: Gross Primary Productivity
- NPP: Net Primary Productivity
- Energy fluxes: W / m2
- Other:
- LAI: m2 / m2
| Variable | Description |
|---|---|
| GPP | Gross Primary Productivity |
| NPP | Net Primary Productivity |
| TotalResp | Total Respiration |
| AutoResp | Autotrophic Respiration |
| HeteroResp | Heterotrophic Respiration |
| SoilResp | Soil Respiration |
| NEE | Net Ecosystem Exchange |
| AbvGrndWood | Above ground woody biomass |
| leaf_carbon_content | Leaf Carbon Content |
| TotLivBiom | Total living biomass |
| TotSoilCarb | Total Soil Carbon |
| Qle | Latent heat |
| Transp | Total transpiration |
| SoilMoist | Average Layer Soil Moisture |
| SoilMoistFrac | Average Layer Fraction of Saturation |
| SWE | Snow Water Equivalent |
| litter_carbon_content | Litter Carbon Content |
| litter_mass_content_of_water | Average layer litter moisture |
| LAI | Leaf Area Index |
| fine_root_carbon_content | Fine Root Carbon Content |
| coarse_root_carbon_content | Coarse Root Carbon Content |
| GWBI | Gross Woody Biomass Increment |
| AGB | Total aboveground biomass |
| time_bounds | history time interval endpoints |
4. Extract SIPNET Output
First, uncompress the model output. Only the netCDF files are needed.
# Set paths
ccmmf_dir=/projectnb2/dietzelab/ccmmf
archive_file="$ccmmf_dir/lebauer_agu_2025_20251210.tgz"
output_dir="$ccmmf_dir/modelout/ccmmf_phase_3_scenarios_20251210"
# Ensure output directory exists
mkdir -p "$output_dir"
#
tar --use-compress-program="pigz -d" -xf \
"$archive_file" \
-C "$output_dir" \
--strip-components=1 \
--no-same-owner #\
# --wildcards \
# 'lebauer_agu_2025/output_/out/' \
# 'lebauer_agu_2025/output_/out/ENS--/[0-9][0-9][0-9][0-9].nc'Rscript scripts/030_extract_sipnet_output.RExtracts and formats SIPNET outputs for downscaling:
- Extract output variables (AGB, TotSoilCarb) from SIPNET simulations
- Aggregate site-level ensemble outputs into long and 4D array formats
- Save CSV and NetCDF files following EFI standards
Inputs: - out/ENS-<ensemble_number>-<site_id>/YYYY.nc - results_monthly_<variable>.Rdata (management scenario mode) - site_info.csv (management scenario mode)
Outputs: - out/ensemble_output.csv: Long format data with columns: - scenario: Management scenario identifier (when scenarios enabled) - datetime, site_id, lat, lon, pft, parameter, variable, prediction
5. Mixed Cropping Systems
Rscript scripts/031_aggregate_sipnet_output.RSimulates mixed-cropping scenarios by combining outputs across two PFTs using combine_mixed_crops() (see Mixed System Prototype). Two methods are supported:
- weighted: area-partitioned mix where
woody_cover + annual_cover = 1 - incremental: preserve woody baseline (
woody_cover = 1) and add the annual delta scaled byannual_cover
combine_mixed_crops() is pool-agnostic: pass any additive quantity expressed per unit area, including instantaneous stocks (kg/m^2) or total flux totals that have already been accumulated over the SIPNET output interval (e.g., hourly or annual kg/m^2 of NEE). (kg/m^2) or total flux over a defined time step.
Outputs include multi_pft_ensemble_output.csv, combined_ensemble_output.csv, and ensemble_output_with_mixed.csv with a synthetic mixed PFT.
6. Downscale, Aggregate to County, and Plot
Rscript scripts/040_downscale.R
Rscript scripts/041_aggregate_to_county.R
Rscript scripts/042_downscale_analysis.R
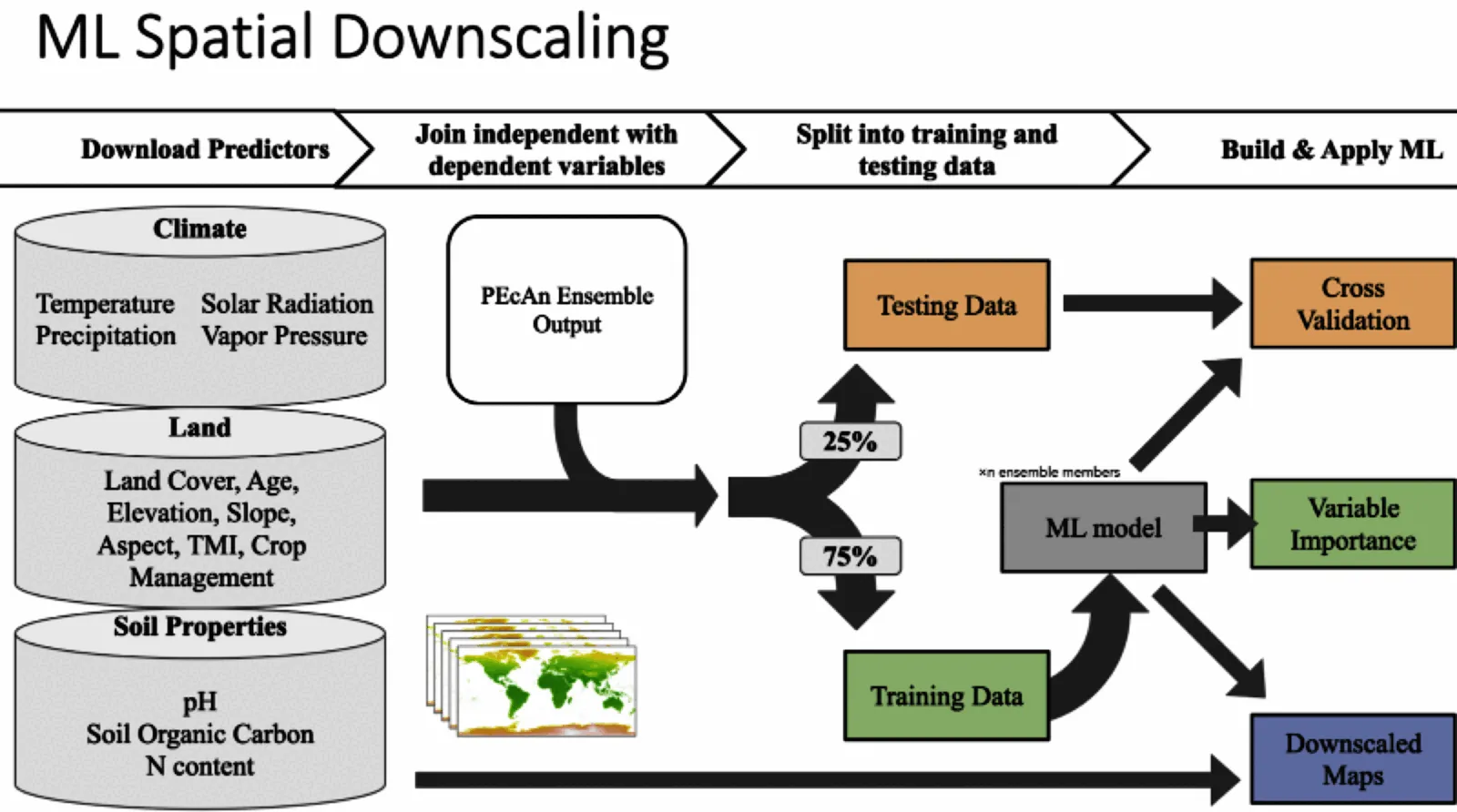
Rscript scripts/043_county_level_plots.RBuilds Random Forest models to predict carbon pools for all fields; aggregates to county-level; summarizes variable importance; and produces maps:
- Train models on SIPNET ensemble runs at design points
- Use environmental covariates to downscale predictions to all fields
- Aggregate to county-level estimates
- Output maps and statistics of carbon density and totals
Inputs: - model_outdir/ensemble_output.csv: SIPNET outputs extracted in step 4 - data/site_covariates.csv: Environmental covariates
Outputs from 040_downscale.R: - model_outdir/downscaled_preds.csv: Per-site, per-ensemble predictions with columns: - site_id, scenario, pft, ensemble, c_density_Mg_ha, total_c_Mg, area_ha, county, model_output - model_outdir/downscaled_preds_metadata.json: Metadata including scenarios list - model_outdir/downscaled_deltas.csv: Carbon change from simulation start to end per scenario - model_outdir/training_sites.csv: Training site list per scenario, PFT, and pool - cache/models/*_models.rds: Saved RF models per spec - cache/training_data/*_training.csv: Training covariate matrices per spec
Outputs from 041_aggregate_to_county.R: - model_outdir/county_aggregated_preds.csv: County statistics per scenario with columns: - scenario, model_output, pft, county, n, mean_total_c_Tg, sd_total_c_Tg, mean_c_density_Mg_ha, sd_c_density_Mg_ha, mean_total_ha, sd_total_ha - model_outdir/county_aggregated_deltas.csv: County-level carbon change per scenario - model_outdir/state_summaries.csv: State-level totals per scenario - model_outdir/aggregation_metadata.json: Metadata for aggregated outputs
Outputs from 042_downscale_analysis.R (saved in figures/): - <pft>_<pool>_importance_partial_plots.png: Variable importance with partial plots for top predictors - <pft>_<pool>_ALE_predictor<i>.svg and <pft>_<pool>_ICE_predictor<i>.svg: ALE and ICE plots
Outputs from 043_county_level_plots.R (saved in figures/): - county_<scenario>_<pft>_<pool>_carbon_stock.webp: County-level stock maps per scenario - field_<scenario>_<pft>_<pool>_carbon_density.webp: Field-level density point maps per scenario - county_diff_woody_plus_annual_minus_woody_<pool>_carbon_stock.webp: Difference maps (mixed - woody) - stock only - county_delta_<scenario>_<pft>_<pool>_carbon_stock.webp: Start-to-end delta stock maps when available
Running on BU Cluster
Interactive session (example):
qrsh -l h_rt=3:00:00 -pe omp 16 -l buyinSubmit a batch job (example using downscaling step):
qsub \
-l h_rt=4:00:00 \
-pe omp 8 \
-o logs/040.out \
-e logs/040.err \
-b y Rscript scripts/040_downscale.RDocumentation Site (Quarto)
- Preview:
quarto preview - Build:
quarto render - Publish to GitHub Pages:
quarto publish gh-pages(publishes_site/)
Notes: - Quarto config is in _quarto.yml and the home page is index.qmd (includes README.md). - Uses freeze: auto to cache outputs; rebuild where data paths exist (see 000-config.R).
Technical Reference
Ensemble Structure
Each ensemble member represents a plausible realization given parameter and meteorological uncertainty. This ensemble structure is maintained throughout the workflow to properly propagate uncertainty. For example, downscaling is done for each ensemble member separately, and then the results are aggregated to county-level statistics.