Model Drivers
Environmental Predictors Driving Spatial Variation in Downscaled Predictions
The Random Forest downscaling model learns relationships between environmental site characteristics and SIPNET model outputs at 100 anchor sites, then predicts to all ~132,000 cropland fields using eight predictors (seven environmental covariates plus field area). The diagnostic plots below reveal which environmental factors drive the spatial variation seen in the maps. See the Scenario Definition Table for practice parameters and simulation details.
These results are from a proof-of-concept modeling framework that has not been validated against field observations. Interpret all values as illustrative projections, not empirical estimates.
Model Performance Overview
Before interpreting predictor effects, it is important to understand how well each Random Forest model captures spatial variation. Out-of-bag (OOB) R² measures predictive accuracy on held-out training sites.
| Variable | Median OOB R² | IQR | Top Predictors | Assessment |
|---|---|---|---|---|
| N2O Flux | 0.52 | 0.38–0.72 | vapr, temp | Best spatial model |
| TotSoilCarb | 0.34 | 0.26–0.41 | ocd, vapr | Moderate |
| AGB | 0.09 | -0.04–0.52 | precip, vapr | Poor – high variance across ensembles |
| CH4 Flux | -0.15 | -0.17–-0.13 | none meaningful | No spatial signal (near-zero baseline) |
Non-flooded annual croplands produce near-zero CH4 fluxes in SIPNET (approximately 10⁻¹⁴ kg m⁻² at the modeled sites) because methanogenesis requires sustained anaerobic conditions (continuous flooding) that do not occur under standard irrigation. With essentially no spatial variation in the training data, the Random Forest has no signal to learn – the negative OOB R² (-0.15) confirms the model performs worse than a constant mean. The CH4 ALE/ICE plots below reflect noise, not ecology.
Environmental Predictors
The downscaling model uses eight predictors (seven environmental covariates plus field area):
| Variable | Description | Source | Units | Ecological Role |
|---|---|---|---|---|
clay |
Clay content | SoilGrids | % | Soil texture controls water holding capacity and physical protection of organic matter |
ocd |
Organic carbon density | SoilGrids | hg/m3 | Existing soil organic matter – both a predictor and correlated with the output |
twi |
Topographic wetness index | SRTM-derived | - | Topographic position controls water accumulation and drainage |
temp |
Mean annual temperature | ERA5 | °C | Temperature drives decomposition rates and plant productivity |
precip |
Mean annual precipitation | ERA5 | mm/yr | Moisture availability for plant growth and microbial activity |
srad |
Solar radiation | ERA5 | J/m² | Energy input for photosynthesis |
vapr |
Vapor pressure | ERA5 | kPa | Atmospheric moisture – strongly correlated with temperature (r = 0.86) |
area_ha |
Field area | CADWR | ha | Structural covariate capturing field-size effects on model predictions |
ALE and ICE Diagnostic Plots
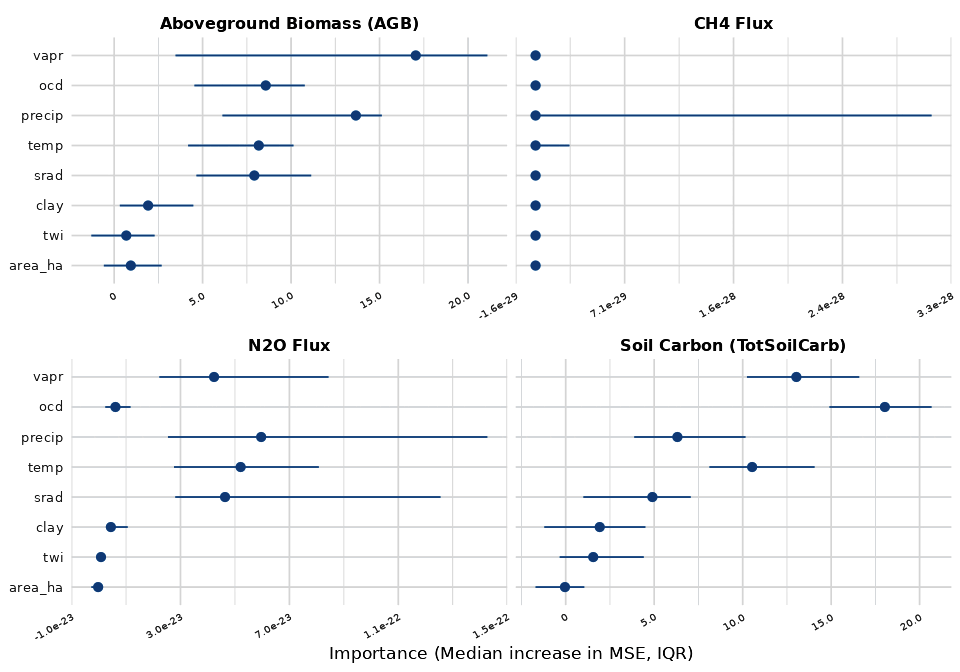
ALE (Accumulated Local Effects) plots show the average marginal effect of each predictor on the model output across its range. Steeper slopes indicate stronger influence.
ICE (Individual Conditional Expectation) plots show how each individual anchor site responds to changes in a predictor while holding all others constant. When ICE lines diverge, it indicates interaction effects – the predictor’s influence depends on other site characteristics.
Total Soil Carbon (TotSoilCarb)
Higher organic carbon density (ocd) –> higher soil carbon: sites with more existing organic matter retain more carbon through the simulation. Vapor pressure (vapr) shows a negative relationship – higher vapr (warmer, more humid conditions) correlates with lower soil carbon, consistent with faster decomposition at higher temperatures (vapr and temp are highly correlated, r = 0.86).
Aboveground Biomass (AGB)
Precipitation (precip) and vapor pressure (vapr) are the top drivers of AGB variation, though this model has low explanatory power (median OOB R² = 0.09). Higher precipitation –> more water availability –> more growth. Vapor pressure is strongly correlated with temperature (r = 0.86); its negative relationship with AGB likely reflects faster turnover under warmer conditions rather than a direct moisture effect. The wide IQR (-0.04 to 0.52) across ensembles indicates the spatial pattern is not reliable.
N2O Flux
Vapor pressure (vapr) and mean annual temperature (temp) are the top predictors of spatial variation in N2O flux (OOB R² = 0.52, our best spatial model). Higher vapr and temperature are associated with greater N2O emissions, consistent with warmer conditions promoting nitrification and denitrification. These two predictors are highly correlated (r = 0.86), reflecting California’s north-south climate gradient.
CH4 Flux
The CH4 downscaling model has negative OOB R² (-0.15), indicating it has no predictive skill for spatial variation. Non-flooded annual croplands produce near-zero CH4 fluxes (raw SIPNET output on the order of 10⁻¹⁴ kg m⁻²; statewide downscaled total roughly 13,600 kg CH4 yr⁻¹) because methanogenesis requires sustained soil saturation that does not occur under standard irrigation, leaving the Random Forest with no spatial signal to learn. The top “predictors” (area_ha, clay) carry no ecological meaning here – they reflect noise fitting. Interpret these plots accordingly.
Why These Drivers Matter
Understanding what drives spatial variation is essential for targeting conservation practices:
- If soil type dominates (clay, ocd) – practice benefits are site-specific, and adoption should prioritize fields with responsive soil types.
- If climate dominates (temp, vapr, precip) – practice effects vary with regional climate, suggesting geographic targeting based on climate zones.
- If the practice effect is uniform regardless of drivers – broad-scale adoption is efficient because benefits are not geographically restricted.
The field-level difference maps in the Soil Carbon Atlas and GHG Emissions Atlas show which pattern applies for each variable and scenario.